Glance accepts a model object and returns a tibble::tibble()
with exactly one row of model summaries. The summaries are typically
goodness of fit measures, p-values for hypothesis tests on residuals,
or model convergence information.
Glance never returns information from the original call to the modeling function. This includes the name of the modeling function or any arguments passed to the modeling function.
Glance does not calculate summary measures. Rather, it farms out these
computations to appropriate methods and gathers the results together.
Sometimes a goodness of fit measure will be undefined. In these cases
the measure will be reported as NA.
Glance returns the same number of columns regardless of whether the
model matrix is rank-deficient or not. If so, entries in columns
that no longer have a well-defined value are filled in with an NA
of the appropriate type.
# S3 method for class 'coxph'
glance(x, ...)Arguments
- x
A
coxphobject returned fromsurvival::coxph().- ...
For
tidy(), additional arguments passed tosummary(x, ...). Otherwise ignored.
See also
Other coxph tidiers:
augment.coxph(),
tidy.coxph()
Other survival tidiers:
augment.coxph(),
augment.survreg(),
glance.aareg(),
glance.cch(),
glance.pyears(),
glance.survdiff(),
glance.survexp(),
glance.survfit(),
glance.survreg(),
tidy.aareg(),
tidy.cch(),
tidy.coxph(),
tidy.pyears(),
tidy.survdiff(),
tidy.survexp(),
tidy.survfit(),
tidy.survreg()
Value
A tibble::tibble() with exactly one row and columns:
- AIC
Akaike's Information Criterion for the model.
- BIC
Bayesian Information Criterion for the model.
- logLik
The log-likelihood of the model. [stats::logLik()] may be a useful reference.
- n
The total number of observations.
- nevent
Number of events.
- nobs
Number of observations used.
See survival::coxph.object for additional column descriptions.
Examples
# load libraries for models and data
library(survival)
# fit model
cfit <- coxph(Surv(time, status) ~ age + sex, lung)
# summarize model fit with tidiers
tidy(cfit)
#> # A tibble: 2 × 5
#> term estimate std.error statistic p.value
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 age 0.0170 0.00922 1.85 0.0646
#> 2 sex -0.513 0.167 -3.06 0.00218
tidy(cfit, exponentiate = TRUE)
#> # A tibble: 2 × 5
#> term estimate std.error statistic p.value
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 age 1.02 0.00922 1.85 0.0646
#> 2 sex 0.599 0.167 -3.06 0.00218
lp <- augment(cfit, lung)
risks <- augment(cfit, lung, type.predict = "risk")
expected <- augment(cfit, lung, type.predict = "expected")
glance(cfit)
#> # A tibble: 1 × 18
#> n nevent statistic.log p.value.log statistic.sc p.value.sc statistic.wald
#> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 228 165 14.1 0.000857 13.7 0.00105 13.5
#> # ℹ 11 more variables: p.value.wald <dbl>, statistic.robust <dbl>,
#> # p.value.robust <dbl>, r.squared <dbl>, r.squared.max <dbl>,
#> # concordance <dbl>, std.error.concordance <dbl>, logLik <dbl>, AIC <dbl>,
#> # BIC <dbl>, nobs <dbl>
# also works on clogit models
resp <- levels(logan$occupation)
n <- nrow(logan)
indx <- rep(1:n, length(resp))
logan2 <- data.frame(
logan[indx, ],
id = indx,
tocc = factor(rep(resp, each = n))
)
logan2$case <- (logan2$occupation == logan2$tocc)
cl <- clogit(case ~ tocc + tocc:education + strata(id), logan2)
tidy(cl)
#> # A tibble: 9 × 5
#> term estimate std.error statistic p.value
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 toccfarm -1.90 1.38 -1.37 1.70e- 1
#> 2 toccoperatives 1.17 0.566 2.06 3.91e- 2
#> 3 toccprofessional -8.10 0.699 -11.6 4.45e-31
#> 4 toccsales -5.03 0.770 -6.53 6.54e-11
#> 5 tocccraftsmen:education -0.332 0.0569 -5.84 5.13e- 9
#> 6 toccfarm:education -0.370 0.116 -3.18 1.47e- 3
#> 7 toccoperatives:education -0.422 0.0584 -7.23 4.98e-13
#> 8 toccprofessional:education 0.278 0.0510 5.45 4.94e- 8
#> 9 toccsales:education NA 0 NA NA
glance(cl)
#> # A tibble: 1 × 18
#> n nevent statistic.log p.value.log statistic.sc p.value.sc statistic.wald
#> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 4190 838 666. 1.90e-138 682. 5.01e-142 414.
#> # ℹ 11 more variables: p.value.wald <dbl>, statistic.robust <dbl>,
#> # p.value.robust <dbl>, r.squared <dbl>, r.squared.max <dbl>,
#> # concordance <dbl>, std.error.concordance <dbl>, logLik <dbl>, AIC <dbl>,
#> # BIC <dbl>, nobs <dbl>
library(ggplot2)
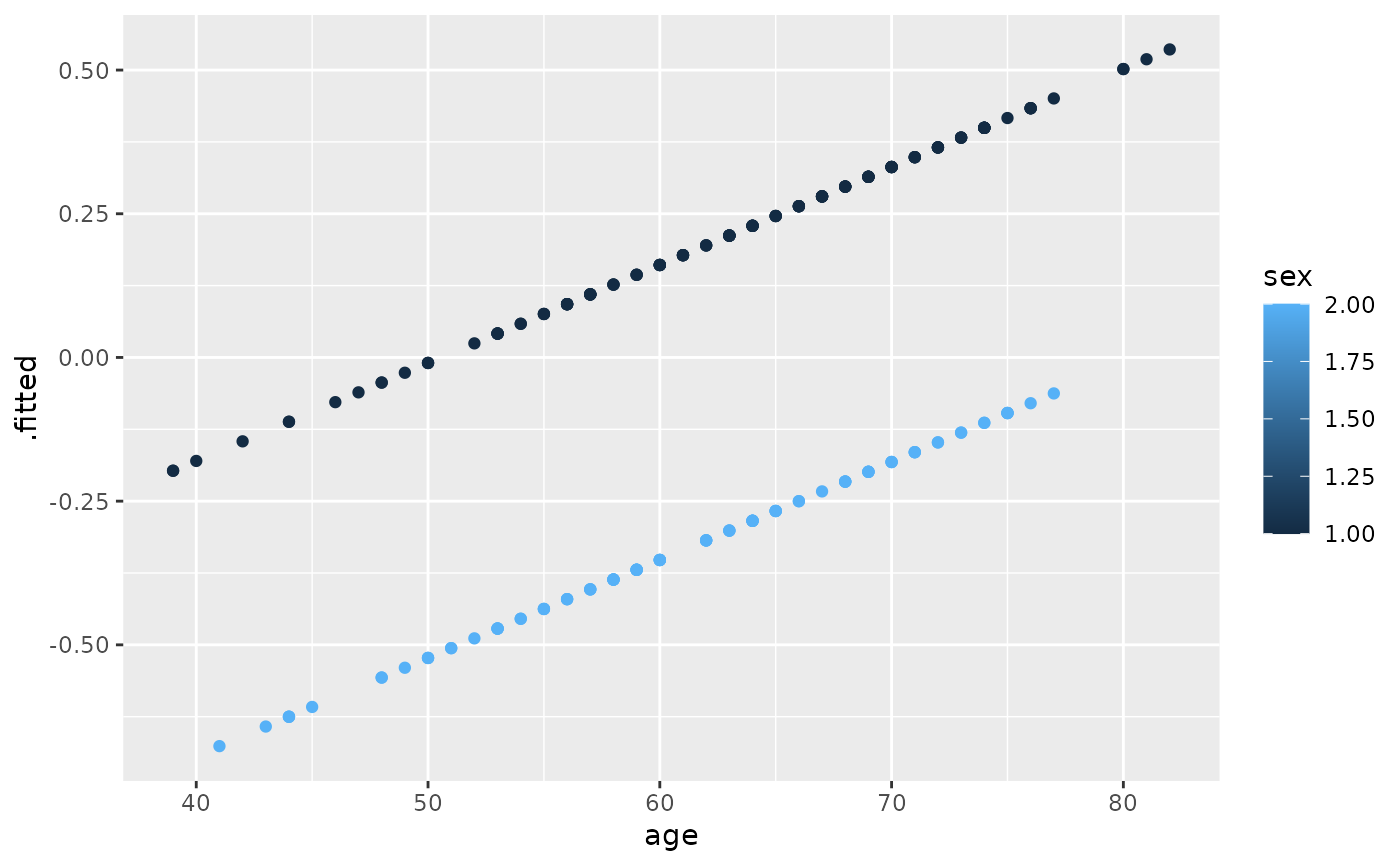
ggplot(lp, aes(age, .fitted, color = sex)) +
geom_point()
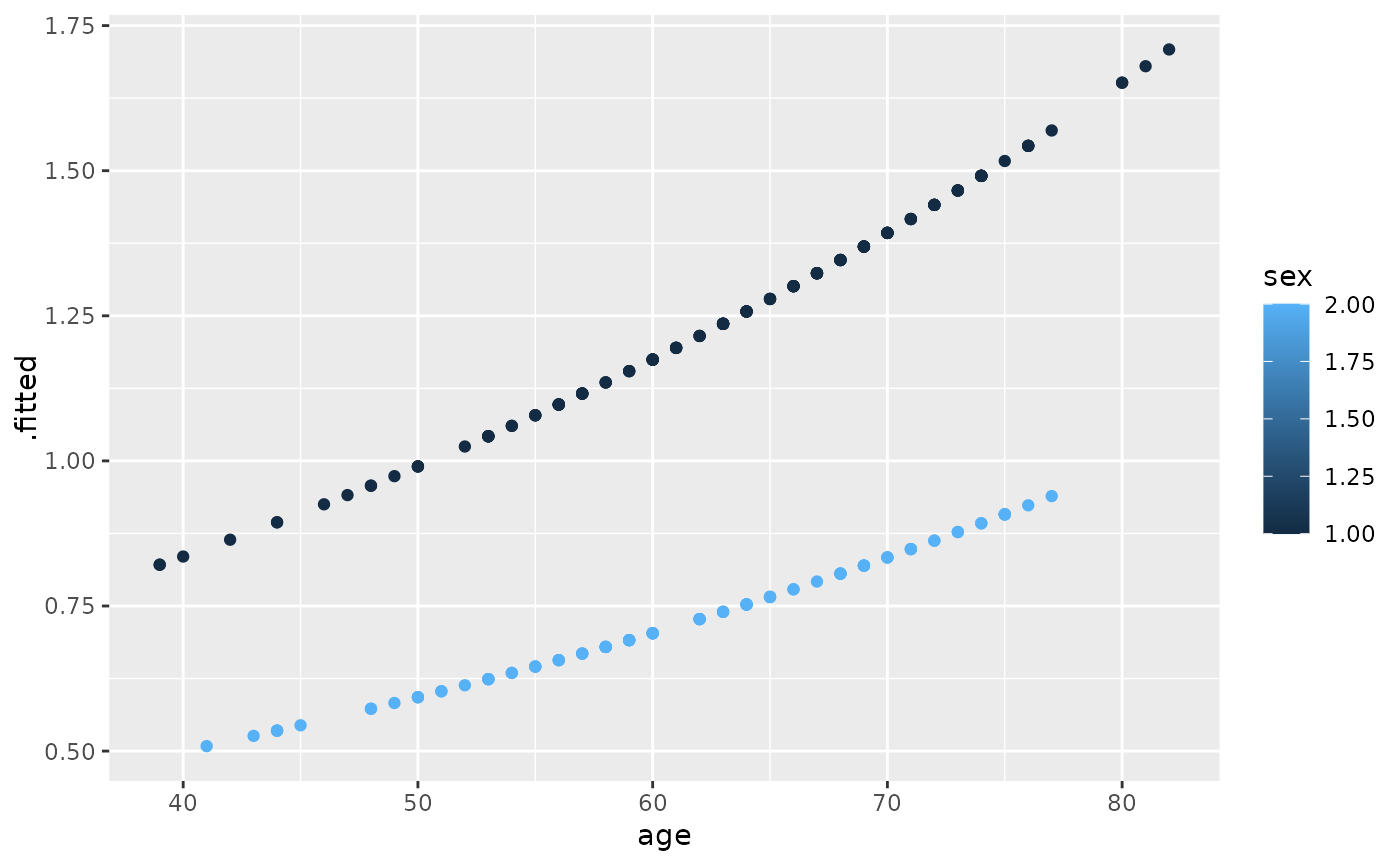
 ggplot(risks, aes(age, .fitted, color = sex)) +
geom_point()
ggplot(risks, aes(age, .fitted, color = sex)) +
geom_point()
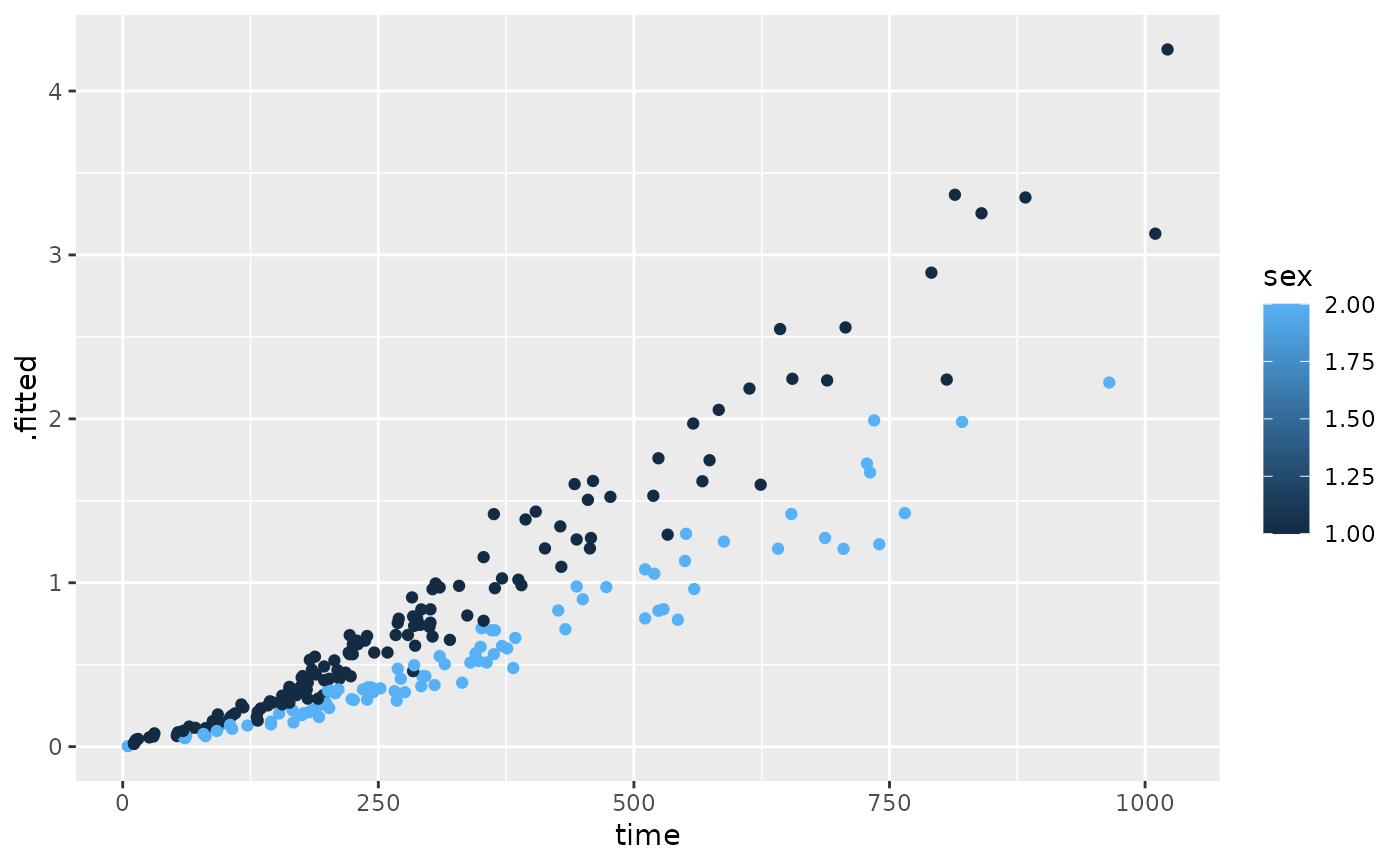
 ggplot(expected, aes(time, .fitted, color = sex)) +
geom_point()
ggplot(expected, aes(time, .fitted, color = sex)) +
geom_point()